Graphormer实战教程:基于ogb库加载PCQM4M数据微调模型示例
news2026/4/1 5:31:00
Graphormer实战教程基于ogb库加载PCQM4M数据微调模型示例1. 引言Graphormer是一种创新的分子属性预测模型采用纯Transformer架构的图神经网络设计。它专门针对分子图原子-键结构的全局结构建模与属性预测任务在OGB、PCQM4M等分子基准测试中表现优异大幅超越传统GNN模型。本教程将带您从零开始学习如何使用ogb库加载PCQM4M数据集并对Graphormer模型进行微调。通过本教程您将掌握如何准备分子图数据环境使用ogb库加载和处理PCQM4M数据集配置和微调Graphormer模型评估模型性能的完整流程2. 环境准备与安装2.1 系统要求Python 3.8CUDA 11.3 (推荐)至少16GB内存支持PyTorch的GPU (推荐RTX 3090及以上)2.2 安装依赖conda create -n graphormer python3.9 conda activate graphormer pip install torch1.12.0cu113 torchvision0.13.0cu113 torchaudio0.12.0 --extra-index-url https://download.pytorch.org/whl/cu113 pip install ogb rdkit-pypi torch-geometric2.3 安装Graphormergit clone https://github.com/microsoft/Graphormer.git cd Graphormer pip install -e .3. 数据准备与处理3.1 了解PCQM4M数据集PCQM4M是OGB(Open Graph Benchmark)提供的大规模分子属性预测数据集包含约380万个分子结构及其HOMO-LUMO能隙值。3.2 加载数据集from ogb.lsc import PCQM4Mv2Dataset dataset PCQM4Mv2Dataset(rootdataset/) print(dataset) print(dataset[0]) # 查看第一个样本3.3 数据预处理Graphormer需要特定的数据格式我们需要将分子图转换为模型可接受的输入from ogb.utils.features import atom_to_feature_vector, bond_to_feature_vector from rdkit import Chem def smiles2graph(smiles_string): mol Chem.MolFromSmiles(smiles_string) # 原子特征 atom_features_list [] for atom in mol.GetAtoms(): atom_features_list.append(atom_to_feature_vector(atom)) # 键特征 edges [] edge_features_list [] for bond in mol.GetBonds(): i bond.GetBeginAtomIdx() j bond.GetEndAtomIdx() edge_feature bond_to_feature_vector(bond) edges.append((i, j)) edge_features_list.append(edge_feature) edges.append((j, i)) # 无向图 edge_features_list.append(edge_feature) # 处理孤立原子 if len(edges) 0: for i in range(len(atom_features_list)): edges.append((i, i)) edge_features_list.append([0]*len(edge_feature)) return atom_features_list, edges, edge_features_list4. 模型配置与微调4.1 加载预训练模型from graphormer import Graphormer model Graphormer( n_layers12, num_heads32, hidden_dim512, dropout_rate0.1, intput_dropout_rate0.1, ffn_dim512, dataset_namepcqm4m, )4.2 数据加载器设置from torch_geometric.data import DataLoader # 划分训练集、验证集 split_idx dataset.get_idx_split() train_dataset dataset[split_idx[train]] valid_dataset dataset[split_idx[valid]] # 创建数据加载器 train_loader DataLoader(train_dataset, batch_size32, shuffleTrue) valid_loader DataLoader(valid_dataset, batch_size32, shuffleFalse)4.3 训练配置import torch import torch.nn as nn from torch.optim import AdamW device torch.device(cuda if torch.cuda.is_available() else cpu) model model.to(device) criterion nn.MSELoss() optimizer AdamW(model.parameters(), lr1e-4, weight_decay0.01) scheduler torch.optim.lr_scheduler.CosineAnnealingLR(optimizer, T_max10)5. 训练与评估5.1 训练循环def train(): model.train() total_loss 0 for batch in train_loader: batch batch.to(device) optimizer.zero_grad() # 前向传播 out model(batch) loss criterion(out, batch.y.view(-1, 1)) # 反向传播 loss.backward() optimizer.step() total_loss loss.item() return total_loss / len(train_loader)5.2 验证循环torch.no_grad() def validate(): model.eval() total_loss 0 for batch in valid_loader: batch batch.to(device) out model(batch) loss criterion(out, batch.y.view(-1, 1)) total_loss loss.item() return total_loss / len(valid_loader)5.3 主训练流程best_val_loss float(inf) patience 5 counter 0 for epoch in range(1, 101): train_loss train() val_loss validate() scheduler.step() print(fEpoch: {epoch:03d}, Train Loss: {train_loss:.4f}, Val Loss: {val_loss:.4f}) if val_loss best_val_loss: best_val_loss val_loss torch.save(model.state_dict(), best_model.pt) counter 0 else: counter 1 if counter patience: print(Early stopping!) break6. 模型评估与应用6.1 测试集评估test_dataset dataset[split_idx[test]] test_loader DataLoader(test_dataset, batch_size32, shuffleFalse) torch.no_grad() def test(): model.eval() total_loss 0 for batch in test_loader: batch batch.to(device) out model(batch) loss criterion(out, batch.y.view(-1, 1)) total_loss loss.item() return total_loss / len(test_loader) test_loss test() print(fTest Loss: {test_loss:.4f})6.2 预测新分子def predict_smiles(smiles): model.eval() atom_features, edges, edge_features smiles2graph(smiles) # 转换为模型输入格式 data { x: torch.tensor(atom_features, dtypetorch.long), edge_index: torch.tensor(edges, dtypetorch.long).t().contiguous(), edge_attr: torch.tensor(edge_features, dtypetorch.long), } data data.to(device) pred model(data) return pred.item() # 示例预测 smiles CCO # 乙醇 prediction predict_smiles(smiles) print(fPredicted HOMO-LUMO gap for {smiles}: {prediction:.4f})7. 总结通过本教程我们完成了以下工作搭建了Graphormer的运行环境并安装了必要依赖使用ogb库加载和处理了PCQM4M数据集配置了Graphormer模型并实现了微调流程评估了模型在验证集和测试集上的性能实现了对新分子属性的预测功能Graphormer作为基于Transformer的图神经网络在分子属性预测任务上展现出强大能力。通过本教程的实践您应该已经掌握了使用ogb库和Graphormer进行分子属性预测的基本流程。获取更多AI镜像想探索更多AI镜像和应用场景访问 CSDN星图镜像广场提供丰富的预置镜像覆盖大模型推理、图像生成、视频生成、模型微调等多个领域支持一键部署。
本文来自互联网用户投稿,该文观点仅代表作者本人,不代表本站立场。本站仅提供信息存储空间服务,不拥有所有权,不承担相关法律责任。如若转载,请注明出处:http://www.coloradmin.cn/o/2470934.html
如若内容造成侵权/违法违规/事实不符,请联系多彩编程网进行投诉反馈,一经查实,立即删除!相关文章
SpringBoot-17-MyBatis动态SQL标签之常用标签
文章目录 1 代码1.1 实体User.java1.2 接口UserMapper.java1.3 映射UserMapper.xml1.3.1 标签if1.3.2 标签if和where1.3.3 标签choose和when和otherwise1.4 UserController.java2 常用动态SQL标签2.1 标签set2.1.1 UserMapper.java2.1.2 UserMapper.xml2.1.3 UserController.ja…
wordpress后台更新后 前端没变化的解决方法
使用siteground主机的wordpress网站,会出现更新了网站内容和修改了php模板文件、js文件、css文件、图片文件后,网站没有变化的情况。
不熟悉siteground主机的新手,遇到这个问题,就很抓狂,明明是哪都没操作错误&#x…
网络编程(Modbus进阶)
思维导图 Modbus RTU(先学一点理论)
概念 Modbus RTU 是工业自动化领域 最广泛应用的串行通信协议,由 Modicon 公司(现施耐德电气)于 1979 年推出。它以 高效率、强健性、易实现的特点成为工业控制系统的通信标准。 包…
UE5 学习系列(二)用户操作界面及介绍
这篇博客是 UE5 学习系列博客的第二篇,在第一篇的基础上展开这篇内容。博客参考的 B 站视频资料和第一篇的链接如下:
【Note】:如果你已经完成安装等操作,可以只执行第一篇博客中 2. 新建一个空白游戏项目 章节操作,重…
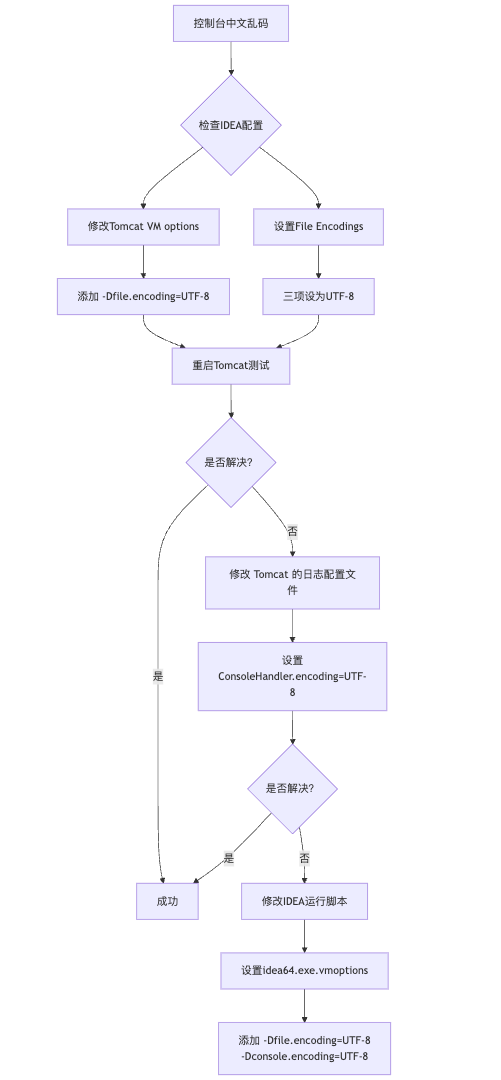
IDEA运行Tomcat出现乱码问题解决汇总
最近正值期末周,有很多同学在写期末Java web作业时,运行tomcat出现乱码问题,经过多次解决与研究,我做了如下整理:
原因:
IDEA本身编码与tomcat的编码与Windows编码不同导致,Windows 系统控制台…
利用最小二乘法找圆心和半径
#include <iostream>
#include <vector>
#include <cmath>
#include <Eigen/Dense> // 需安装Eigen库用于矩阵运算 // 定义点结构
struct Point { double x, y; Point(double x_, double y_) : x(x_), y(y_) {}
}; // 最小二乘法求圆心和半径 …
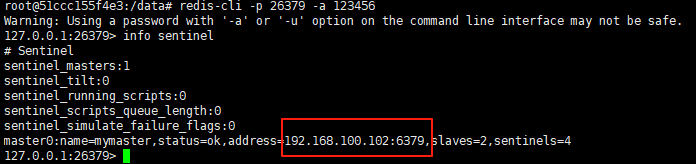
使用docker在3台服务器上搭建基于redis 6.x的一主两从三台均是哨兵模式
一、环境及版本说明
如果服务器已经安装了docker,则忽略此步骤,如果没有安装,则可以按照一下方式安装: 1. 在线安装(有互联网环境): 请看我这篇文章 传送阵>> 点我查看 2. 离线安装(内网环境):请看我这篇文章 传送阵>> 点我查看
说明:假设每台服务器已…
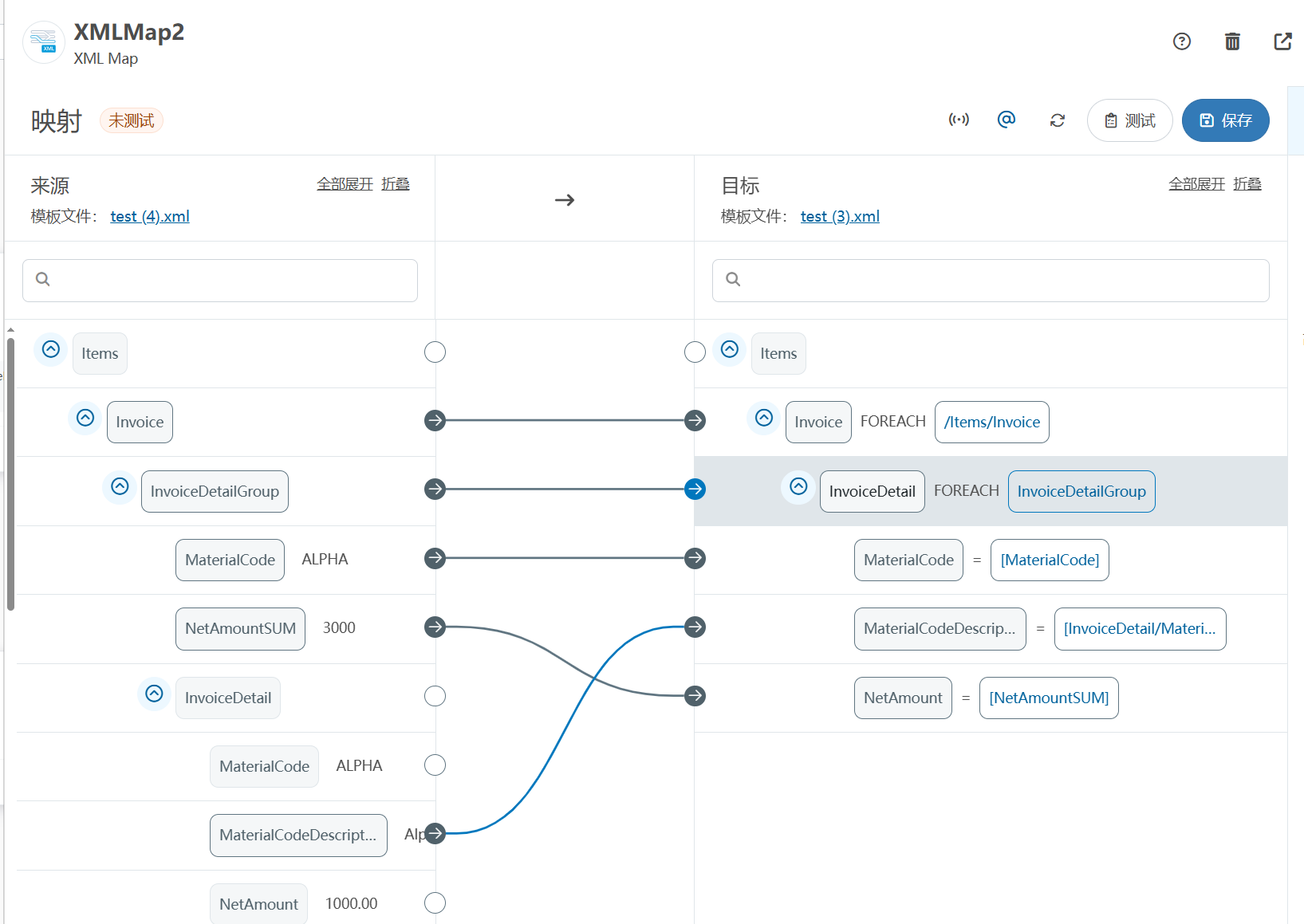
XML Group端口详解
在XML数据映射过程中,经常需要对数据进行分组聚合操作。例如,当处理包含多个物料明细的XML文件时,可能需要将相同物料号的明细归为一组,或对相同物料号的数量进行求和计算。传统实现方式通常需要编写脚本代码,增加了开…
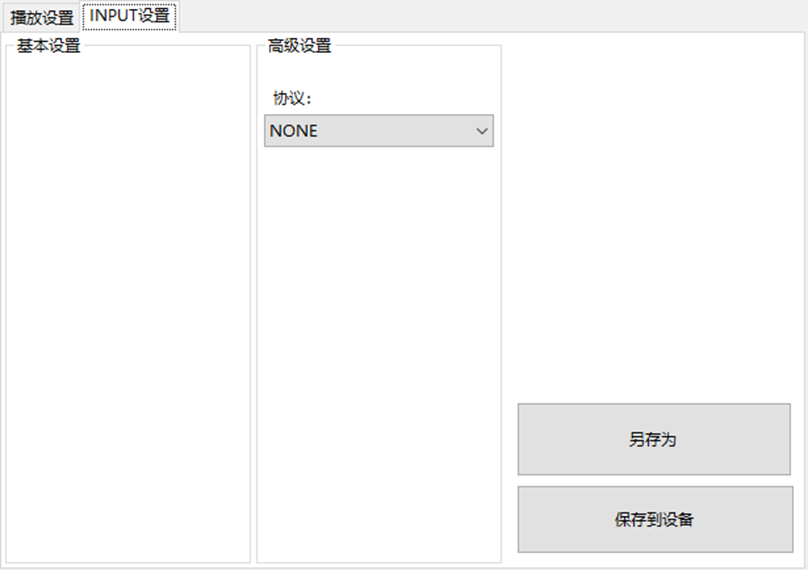
LBE-LEX系列工业语音播放器|预警播报器|喇叭蜂鸣器的上位机配置操作说明
LBE-LEX系列工业语音播放器|预警播报器|喇叭蜂鸣器专为工业环境精心打造,完美适配AGV和无人叉车。同时,集成以太网与语音合成技术,为各类高级系统(如MES、调度系统、库位管理、立库等)提供高效便捷的语音交互体验。
L…
(LeetCode 每日一题) 3442. 奇偶频次间的最大差值 I (哈希、字符串)
题目:3442. 奇偶频次间的最大差值 I 思路 :哈希,时间复杂度0(n)。 用哈希表来记录每个字符串中字符的分布情况,哈希表这里用数组即可实现。
C版本:
class Solution {
public:int maxDifference(string s) {int a[26]…
【大模型RAG】拍照搜题技术架构速览:三层管道、两级检索、兜底大模型
摘要
拍照搜题系统采用“三层管道(多模态 OCR → 语义检索 → 答案渲染)、两级检索(倒排 BM25 向量 HNSW)并以大语言模型兜底”的整体框架: 多模态 OCR 层 将题目图片经过超分、去噪、倾斜校正后,分别用…
【Axure高保真原型】引导弹窗
今天和大家中分享引导弹窗的原型模板,载入页面后,会显示引导弹窗,适用于引导用户使用页面,点击完成后,会显示下一个引导弹窗,直至最后一个引导弹窗完成后进入首页。具体效果可以点击下方视频观看或打开下方…
接口测试中缓存处理策略
在接口测试中,缓存处理策略是一个关键环节,直接影响测试结果的准确性和可靠性。合理的缓存处理策略能够确保测试环境的一致性,避免因缓存数据导致的测试偏差。以下是接口测试中常见的缓存处理策略及其详细说明:
一、缓存处理的核…
龙虎榜——20250610
上证指数放量收阴线,个股多数下跌,盘中受消息影响大幅波动。 深证指数放量收阴线形成顶分型,指数短线有调整的需求,大概需要一两天。 2025年6月10日龙虎榜行业方向分析 1. 金融科技
代表标的:御银股份、雄帝科技
驱动…
观成科技:隐蔽隧道工具Ligolo-ng加密流量分析
1.工具介绍
Ligolo-ng是一款由go编写的高效隧道工具,该工具基于TUN接口实现其功能,利用反向TCP/TLS连接建立一条隐蔽的通信信道,支持使用Let’s Encrypt自动生成证书。Ligolo-ng的通信隐蔽性体现在其支持多种连接方式,适应复杂网…
铭豹扩展坞 USB转网口 突然无法识别解决方法
当 USB 转网口扩展坞在一台笔记本上无法识别,但在其他电脑上正常工作时,问题通常出在笔记本自身或其与扩展坞的兼容性上。以下是系统化的定位思路和排查步骤,帮助你快速找到故障原因:
背景:
一个M-pard(铭豹)扩展坞的网卡突然无法识别了,扩展出来的三个USB接口正常。…
未来机器人的大脑:如何用神经网络模拟器实现更智能的决策?
编辑:陈萍萍的公主一点人工一点智能 未来机器人的大脑:如何用神经网络模拟器实现更智能的决策?RWM通过双自回归机制有效解决了复合误差、部分可观测性和随机动力学等关键挑战,在不依赖领域特定归纳偏见的条件下实现了卓越的预测准…
Linux应用开发之网络套接字编程(实例篇)
服务端与客户端单连接
服务端代码
#include <sys/socket.h>
#include <sys/types.h>
#include <netinet/in.h>
#include <stdio.h>
#include <stdlib.h>
#include <string.h>
#include <arpa/inet.h>
#include <pthread.h>
…
华为云AI开发平台ModelArts
华为云ModelArts:重塑AI开发流程的“智能引擎”与“创新加速器”!
在人工智能浪潮席卷全球的2025年,企业拥抱AI的意愿空前高涨,但技术门槛高、流程复杂、资源投入巨大的现实,却让许多创新构想止步于实验室。数据科学家…
深度学习在微纳光子学中的应用
深度学习在微纳光子学中的主要应用方向
深度学习与微纳光子学的结合主要集中在以下几个方向:
逆向设计 通过神经网络快速预测微纳结构的光学响应,替代传统耗时的数值模拟方法。例如设计超表面、光子晶体等结构。
特征提取与优化 从复杂的光学数据中自…